Discover
Visualize rapid, validated insights through real-world data.
Datavant Completes Acquisition of Aetion. Learn more →

Earlier this year, Aetion co-authored a study published in Science that demonstrates how real-world evidence (RWE) can inform drug development by taking lab-generated hypotheses and assessing them in real clinical practice.
The study, which was led by Nevan Krogan, Ph.D. of the Quantitative Biosciences Institute at the University of California San Francisco along with nearly 200 international collaborators, generated hypotheses that suggested certain medications—those that block the coronavirus’s interaction with the enzyme PGES-2 or sigma receptor 1—may inhibit viral replication of three lethal coronaviruses in vitro. Aetion researchers then used real-world data (RWD) to assess the effectiveness of these medications on COVID-19 severity in real-world patient cohorts (in vivo).
Aetion scientists used Real-Time Insights and Evidence, a COVID-specific instance of the Aetion Evidence Platform®, to run RWE analyses on a select set of the promising drug candidates identified in the lab. Read on to learn more about our work to support this study.
Intent of the research
Aetion’s goal as part of this research project was to evaluate the hypotheses derived from molecular insights in U.S. COVID-19 patients who are treated in real-world settings. To do so, we needed to define a fit-for-purpose, real-world dataset to identify treated patients and controls within the data, match patients on time and other characteristics, and assess whether COVID-19 severity decreased among the treated patients.
Process for the analyses
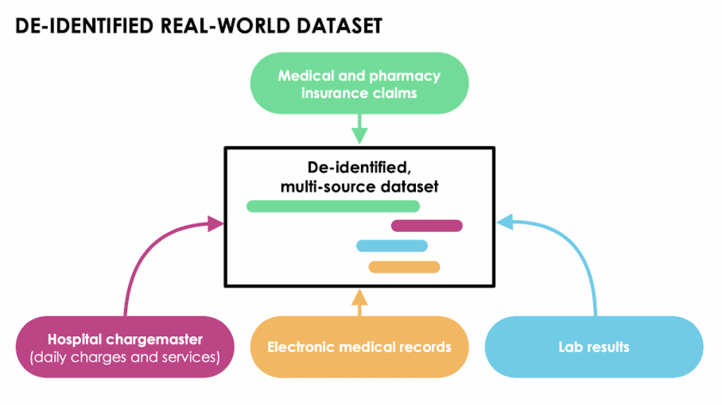
Our first task was to see whether the RWD was fit to answer the research question at hand. This started with understanding whether we could identify patients with COVID-19 who, shortly after their COVID diagnosis, began treatment with one of the potentially effective therapies. We analyzed a linked dataset drawn from medical claims, pharmacy claims, laboratory data, and hospital chargemaster data—which included nearly 740,000 COVID-19 patients—to identify a study population.
We identified COVID-19 patients who had started a course of treatment with one of the two medications determined as promising candidates during the lab portion of the study—indomethacin and typical antipsychotics—and compared them to a group of patients who had started an alternative therapy. We first studied indomethacin, a marketed non-steroidal anti-inflammatory drug (NSAID), in the outpatient setting, and compared it to celecoxib, a newer NSAID. We then looked at the class of typical antipsychotic medications in the inpatient setting compared to atypical antipsychotics.
We then worked to achieve balance between the study groups at baseline, similar to the balancing achieved by randomization in a randomized trial. To do this, we relied on two techniques, risk-set sampling to select comparable referent patients for each treated patient, and propensity score matching to remove any additional imbalances. We then followed each group for markers of worsening severity of COVID-19, and made the comparison between the medications.
Findings from Aetion’s analyses
Aetion’s analyses reached two major findings. First, though the confidence interval was wide, COVID-19-postive users of indomethacin—which targets the PGES-2 enzyme—appeared less likely to require hospitalization or inpatient services than matched users of celecoxib—which doesn’t target PGES-2.
In addition, hospitalized COVID-19-positive new users of typical antipsychotics that act against the sigma receptor 1 protein—such as haloperidol or chlorpromazine—were over 50 percent less likely to require mechanical ventilation compared to matched users of atypical antipsychotics like quetiapine or olanzapine.

Typical antipsychotics can have significant adverse effects, which would disfavor their prophylactic or therapeutic use for patients with COVID-19. That said, these results support further investigation into other medications that target the sigma receptor 1 protein—such as fluvoxamine. In addition, given the high prevalence of neurological complications among hospitalized COVID-19 patients, further validation of sigma receptor findings may support clinicians in a choice between typical or atypical antipsychotic medications.

Implications of the research
This study provides a novel example of how RWE can accelerate the clinical research process by assessing molecular insights to help to identify drug repurposing candidates, or to inform development of a new medication by assessing existing drugs with similar mechanisms of action. If other researchers are able to validate these findings over time, they could inform clinical practice and regulatory decision-making for COVID-19 treatment, or use similar methods to support research in other disease areas.
The work also highlights the usefulness of RWE platforms, which support rapid and iterative RWD analyses, to perform such analyses at the highest levels of scientific rigor. Aetion Evidence Platform allowed us to go from hypothesis to results in a matter of weeks—which is critical given the reach and severity of this pandemic.
Ultimately, as COVID-19 treatment candidates emerge, RWE generated according to principled database epidemiology has an important role to play in complementing clinical trials with real-world insights on usage, safety, and effectiveness across the drug lifecycle.